How to set up FIX on your system
About this guide
This post goes through some common issues with setting up FIX. I’ve kept it short since the FIX installation guide already is pretty exhaustive. Nevertheless, I have spelled out some of the steps in the guide more explicitly here and included fixes for a few issues that were not covered.
Installing FIX
Start by downloading and installing FIX as described in the FIX installation guide. Then go through the steps in the FIX README to complete the setup.
Matlab Compilation
If you have a matlab license, the easiest method to set up FIX on your server is to just run the build_MATLAB script in the FIX folder as described in the FIX README. It’s not the preferred method but was the easiest to debug for me. I would recommend this method only for setting up FIX on University servers that are guaranteed to have continuous access to a MATLAB license. The build_MATLAB should run without errors, apart from the last step, which compiles a script for writing CIFTIs as part of the HCP pipeline. For the standard FIX usage, you don’t need those scripts, so you can move on. If you do want those functions, have a look at the README that can tell you how to set up your settings.sh file so that these functions can compile too without error.
Setting up Matlab for FIX
If you run into an error that looks like this:
Error using call_fsl (line 36)
FSL call (/bin/sh -c '. /software/fsl/6.0.4/etc/fslconf/fsl.sh; fslstats /srv/diedrichsen/data/FunctionalFusion/../Cerebellum/super_cerebellum/resting_state/imaging_data/s02/run01_smoothed.ica/FIX/dummy -S > /srv/diedrichsen/data/FunctionalFusion/../Cerebellum/super_cerebellum/resting_state/imaging_data/s02/run01_smoothed.ica/FIX/dummy1.txt') failed, /bin/sh: 1: cannot create /srv/diedrichsen/data/FunctionalFusion/../Cerebellum/super_cerebellum/resting_state/imaging_data/s02/run01_smoothed.ica/FIX/dummy1.txt: Directory nonexistent
Then your FIX installation is likely having trouble running fsl commands from within matlab. But to verify that this is the problem, run the command matlab is trying to run from your terminal, in this case fslstats /srv/diedrichsen/data/FunctionalFusion/../Cerebellum/super_cerebellum/resting_state/imaging_data/s02/run01_smoothed.ica/FIX/dummy -S > /srv/diedrichsen/data/FunctionalFusion/../Cerebellum/super_cerebellum/resting_state/imaging_data/s02/run01_smoothed.ica/FIX/dummy1.txt
. If this produces the expected output, the problem is with running FSL from matlab. In order to make sure you can run FSL commands from within matlab, ammend your startup file to use the FSL setup commands. For that, open matlab and run the command edit(fullfile(userpath,'startup.m')) (as described here). This will create a startup file within your matlab user folder (usually ~/Documents/MATLAB/) if it doesn’t exist already, and open the file in a text editor. Add the following to your startup file:
% FSL Setup
setenv( 'FSLDIR', '/usr/local/fsl' );
setenv('FSLOUTPUTTYPE', 'NIFTI_GZ');
fsldir = getenv('FSLDIR');
fsldirmpath = sprintf('%s/etc/matlab',fsldir);
path(path, fsldirmpath);
clear fsldir fsldirmpath;
Replace /usr/local/fsl with the path to your fsl installation. For the CBS Server, this is /srv/software/fsl/6.0.4. If you inspect your normal FSLDIR (run echo $FSLDIR in your terminal) you will see that this usually points to /software/fsl/6.0.4, instead of pointing to the server installation. You can also use this FSLDIR, but importantly, you will then have to adjust the call_fsl.m script in the $FSLDIR/etc/matlab folder to comment in lines 14 & 23 which enable you to call fsl from matlab on a Ubuntu/Debian system (which the CBS server has). If you don’t do this, then FIX will fail at the extraction stage where it’s trying to run fsl commands from Matlab. I managed to get write permissions for the /srv/software/fsl/6.0.4/etc/matlab/call_fsl.m script and commented in those needed lines. So unless you have a strong reason to use the server’s FSL folder, then use this. You don’t have to use the server’s FSL folder for anything else, you can leave your normal FSLDIR pointing still to /software/fsl/6.0.4 .
Edit 02.09.2024: Be careful about what else you put into your startup.m file. I accidentally added the path to a conflicting matlab version to my startup.m to run SPM code. It gave me the same error I encountered before (
.../dummy1.txt: Directory nonexistent) even though my startup.m file still included the FSL FIX environment setup code. Since I will need to run both SPM and FIX with this startup.m file, I found a somewhat hacky workaround: I added an if-clause that runs different sections of the startup.m file, depending on whether matlab will be called from the command line (as done by FIX) or through the IDE (as I do when running SPM). Now FIX can correctly call FSL again without running into environment issues. However, this is not very elegant and very error-prone, for example if I forget about this arrangement and try to call SPM from the command line (bound to happen in ~6 months). If you have a better solution, feel free to email me and I will add it to the blog post! My current solution looks like this:
% startup.m
if usejava('desktop')
% Code for when MATLAB is launched from the IDE
fprintf('MATLAB started from the IDE.\n');
% Add IDE-specific startup configurations here
else
% Code for when MATLAB is launched from the command line
fprintf('MATLAB started from the command line.\n');
% Add command line-specific startup configurations here
end
Install R
If not installed already on your server, install R as described in the README, which also specifies the necessary packages.
Set R library path to a folder you have write permissions for
On the CBS server there is an R installation available, just run R from your terminal (note the capitalization) to start R. Make sure that the R installation called form the terminal points to your local installation by running which R in your terminal, so that you can install libraries for this R installation and not run into missing write permissions. If the R installation doesn’t point to the local installation then change your profile so that it does. Also be sure to check the FSL_FIX_R_CMD variable in the settings.sh file in the FIX folder. Change the variable to your local R installation (in my case that’s /usr/bin/R).
Set R library path to a folder you have write permissions for
To make sure you install packages into your local R library, set the R_LIBS environment variable to your local R library folder. Put this into your shell profile, so that this is set automatically:
# R Setup for FIX
echo "export R_LIBS=/home/ROBARTS/cnetteko/R" >> ~/.profile
Also make sure that all other R libraries are removed, so that there are no conflicting packages:
# Remove all other R libraries
echo 'export R_LIBS_SITE=" "' >> ~/.profile
My profile file is named ‘.profile’, but check what your profile is named or whether exists at all in your home directory. Possible names for the existing profile file are ‘.zshrc’ ‘.bashrc’, ‘.zprofile’, ‘.bash_profile’ or ‘.profile’.
Once you’ve set the path, open a new terminal and check that the R path is now showing to the correct directory:
# R Setup for FIX
echo $R_LIBS
Now you should be able to install your R packages.
Install R packages
Contrary to what it says in the README, the syntax for R has changed slightly for newer versions, so run:
install.packages("kernlab", version="0.9-24")
install.packages("ROCR", version="1.0-7")
install.packages("class", version="7.3-14")
install.packages("mvtnorm", version="1.0.8")
install.packages("multcomp", version="1.4-8")
install.packages("sandwich") # Before installing "coin" and "party", I had to also install the package "sandwich" - this was not specified in the FIX README
install.packages("gplots",dependencies = TRUE) # not specified in the FIX README, but needed by "ROCR"
install.packages("coin", version="1.2.2")
install.packages("party", version="1.0-25",dependencies = TRUE)
install.packages("e1071", version="1.6-7")
install.packages("randomForest", version="4.6-12")
Troubleshooting R Package installation
randomForest version not available
You might run into an error saying that the a specific package version (e.g. randomForest Version 4.6-12) is not available for this R version. To still install this specific version of the package in R, first, visit the CRAN Archive to choose the version. Right-click on the version and select “Copy Link Location.” Then, store the link location in a variable named urlPackage in R. For example:
urlPackage <- "https://cran.r-project.org/src/contrib/Archive/randomForest/randomForest_4.6-12.tar.gz"
install.packages(urlPackage, repos=NULL, type="source")
Coin and party package troubleshooting
For the ‘coin’ and ‘party’ package, things can unfortunately be a bit trickier. I had to apply this solution to get the correct coin package version installed. Run from within R:
install.packages("devtools", lib="~/R/x86_64-pc-linux-gnu-library/3.6")
install.packages("coin",lib="~/R/x86_64-pc-linux-gnu-library/3.6", dependencies=TRUE)
remove.packages("coin",lib="~/R/x86_64-pc-linux-gnu-library/3.6")
Then run from the terminal command line:
wget https://cran.r-project.org/src/contrib/Archive/coin/coin_1.2-2.tar.gz
R CMD INSTALL coin_1.2-2.tar.gz --library="~/R/x86_64-pc-linux-gnu-library/3.6"
urlPackage <- "https://cran.r-project.org/src/contrib/Archive/party/party_1.0-25.tar.gz"
install.packages(urlPackage, repos = NULL, type = "source")
library("party")
Note that I changed the R CMD INSTALL lib option to –library, since otherwise you get a permission error.
If R is not properly installed or one of the necessary packages is faulty or missing, there will unfortunately be no interpretable output message from FIX. But hints are that there are no .RData files generated and that the call_matlab.sh.o* file contains the error “tail: cannot open `subjectID/xx.feat//fix4melview_training_LOO_thr1.txt' for reading: No such file or directory”. That means the fix4melview files have not been created - likely because R couldn’t run (However, this could also indicate another issue relating to the hand_labels_noise.txt files).
To understand what’s going on, look into the fix-generated log file in one of your subject folders (cat /resting_state/s03/run01_smoothed.ica/.fix_2b_predict.log), check out the FIX FAQ and look into the logMATLAB.txt file (cat /resting_state/s03/run01_smoothed.ica/fix/logMatlab.txt) and the call_matlab.sh.o* file (cat fix_output_folder/call_matlab.sh.o1590473). Checking the paths for your R installation and the availability of the necessary packages from the R command line will also help you figure out what the source of the problem is.
Check R Package installation
Check that you have the correct versions installed by running:
packageVersion('kernlab') # fix version="0.9-24"
packageVersion('ROCR') # fix version="1.0-7"
packageVersion('class') # fix version="7.3-14"
packageVersion('mvtnorm') # fix version="1.0.8" # My version: 1.1.1
packageVersion('multcomp') # fix version="1.4-8" # My version: 1.4.13
packageVersion('coin') # fix version="1.2.2" # My version: 1.3.1
packageVersion('party') # fix version="1.0-25"
packageVersion('e1071') # fix version="1.6-7" # My version: 1.7.14
packageVersion('randomForest') # fix version="4.6-12"
Also check that all libraries can be loaded without error by running:
library('kernlab')
library('ROCR')
library('class')
library('mvtnorm')
library('multcomp')
library('coin')
library('party')
library('e1071')
library('randomForest')
If these all load without error, you’re good to go!
Troubleshooting
Change to Pyfix
Since FSL release 6.0.7.8, FIX uses the python-based implementation pyfix. If you attempt to use old training weights (generated with FSL before version 6.0.7.8), it will throw an error like this:
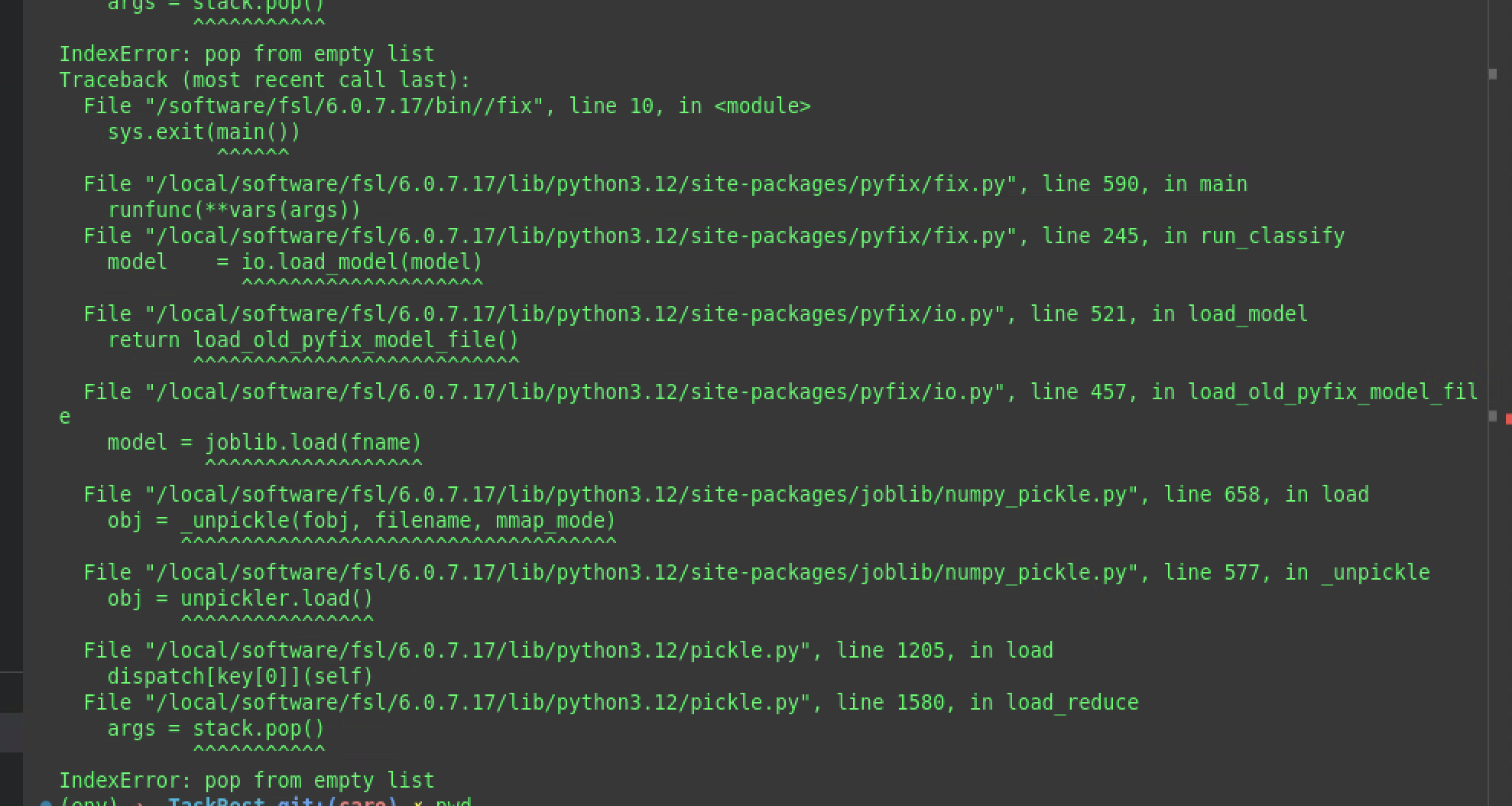
You can update your old training weights using the convert_fix_model function that comes with the new FSL version. Have a look here.